
Cellular Thermal Shift Assay CETSA
One assay,
endless possibilities
What is CETSA?
CETSA®(Cellular Thermal Shift Assay) is our patented core technology and is a cell-based approach that measures shifts in a protein’s thermal stability when a compound binds to it. As the temperature rises, proteins unfold. Binding can stabilize a target and shift its apparent melting behavior. In CETSA, samples are heated across a gradient, and the remaining soluble protein is quantified. This yields a thermal profile that reflects engagement. Results come from temperature-dependent denaturation curves read by immunodetection or mass spectrometry, without engineering tags or overexpressing proteins. This can all be summarized in four simple steps:
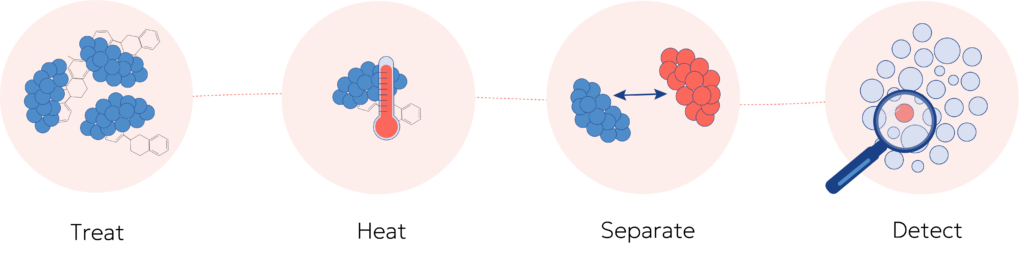
CETSA at a glance
Real biology
Measures compound–target binding in intact cells or lysates
Label and tag-free
Measures native targets-no tags or overexpression needed
One solution for every stage
Supports decisions from target discovery to preclinical studies
CETSA can be applied in every step of the drug discovery value chain, and no matter which cellular matrix you choose, we generate biologically relevant data that helps you prioritize and confirm drug candidates faster.
Intellectual Property
Pelago Bioscience was founded to provide and develop the patented Cellular Thermal Shift Assay (CETSA) in 2013. The CETSA method is patented in most regions worldwide (IP documentation).
Read the paper that introduced CETSA
The original CETSA study introducing the approach and that lead to the founding of Pelago Bioscience: Science, 2013
See CETSA in action
See examples of how CETSA can be used to progress drug discovery projects

Why cellular target engagement matters
Many early choices are made on purified systems that do not reflect real cellular conditions. CETSA brings cell context into those choices. Teams use it to:
• Confirm that hits reach and bind the intended target in a cellular environment
• Rank compounds within a chemotype using comparable engagement metrics
• Support mechanism-of-action claims with evidence from native systems
• Avoid advancing compounds that look strong in vitro but show weak engagement in cells
Where CETSA fits in the pipeline
Discovery: See how proteins behave in real cells, and turn complex biology into clear, actionable insights.
Lead generation: Confirm engagement for prioritized hits and guide chemistry toward viable series
Lead optimization: Track potency shifts and structure–activity trends using thermal profiles
Preclinical research: Examine selectivity and potential off-targets in complex samples

Let’s talk
For us, every project is a partnership.
Send us a message or book a free scoping call with one of our experts.
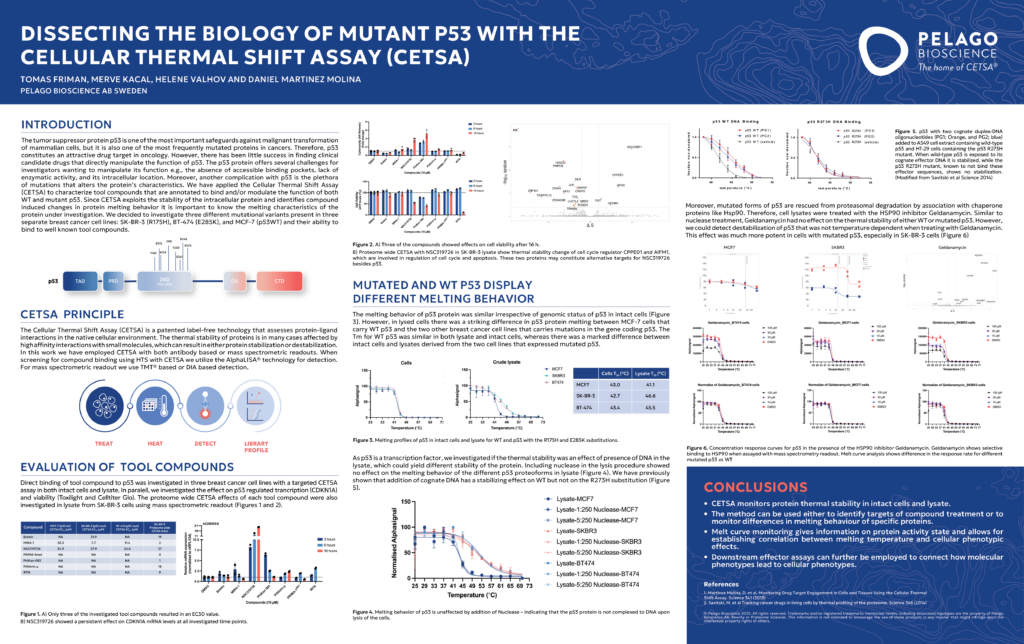
Dissecting the Biology of Mutant p53 with CETSA®
Unlocking the Complex Biology of Mutant p53 with CETSA: Revealing Target Engagement and Stability Profiles p53 target engagement is a critical factor in modern cancer drug discovery. The tumor suppres…

Protein Degradation – Are Your Degraders Delivering?
Uncover Mechanisms with CETSA® This application note explores how CETSA reveals target specificity and mechanisms of action, helping optimize degrader molecules for clinical success. Learn from …

CETSA: Your Key to Biological Relevance
Unlock Deeper Insights in Drug Discovery with CETSA Discover how to accelerate your drug discovery process with Pelago Bioscience’s exclusive eBook, CETSA: Your Key to Biological Relevance…
